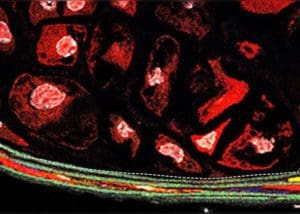
In aortic aneurysm, smooth muscles transform, causing lesions. Photo courtesy of YaleNews.
Researchers at privately-held Genuity Science and Yale University Medical School have revealed why aortic aneurysms form. The scientists discovered that an expansion in abnormal cells in smooth muscle tissue of the aorta can contribute to aortic aneurysm.
The researchers came to that conclusion by using a combination of generative artificial intelligence, single-cell biology and RNA sequencing to uncover causal molecular drivers of cell fate transition in aortic aneurysms.
Subsequent investigations have revealed a possible genetic cause of the disease in humans, by specifically linking a genetic variant in a gene of unknown significance to disease pathogenesis.
In 2018, almost 10,000 people in the U.S. died from ruptured aortic aneurysms, according to the CDC. Currently, the first line of defense against the disease is surgical intervention. These findings could potentially lead to new drug targets and therapies for aneurysms.
The researchers published their findings in Cell Stem Cell.
Uncovering molecular rules that govern cellular behavior
Although the research drew on various disciplines, generative AI was at the heart of the breakthrough. AI is “now capable of teaching us the molecular rules that govern cellular behavior,” according to Tom Chittenden, chief data science officer at Boston-based Genuity Science.
Using a combination of data science techniques, the Genuity team analyzed single-cell RNA sequencing (scRNA-seq) data to identify an aberrant cell cluster resulting from reprogrammed smooth muscle cells. “We were able to identify a cell population that was driving thoracic aortic aneurysm,” Chittenden said.
Tracking global gene expression profiles over a four-month period, the group discovered the loss of expression of a number of smooth muscle cell markers, including myosin heavy chain 11 (MYH11). In the same cell populations where the expression of smooth muscle cell markers were lost, the research team found increased expression of other genes associated with both mesenchymal-like stem cells that ultimately give rise to adipocytes, chondrocytes, and osteoblasts, as well as macrophage-like cells.
“I’ve never seen such a degree of linked causal molecular dysregulation before,” Chittenden said, referring to the genomic profiling changes.
“These cells were dramatically transforming, they were becoming more inflammatory” Chittenden added.
The loss of normal aortic smooth muscle cells and the resulting increase in mass of other tissues ultimately causes a series of changes. In addition to increased inflammation, the reprogrammed smooth muscle cells lead to the growth and dilation of the aorta and the calcification and ossification of the aortic wall. Those changes, in turn, cause aortic aneurysms, the researchers concluded.
To validate the scRNA-seq findings, Yale University Medical School professor Dr. Michael Simons led a team to use a number of additional experimental techniques, including imaging mass cytometry, to identify and characterize a single aberrant cluster of abnormal smooth muscle cells piercing the endothelium of the aorta.
The collaboration with professor Simons and his team afforded even greater insights into disease etiology. “The proteomic markers allowed us to quantify pathologic processes associated with atherosclerosis such as calcification and ossification, which often lead to plaque rupture,” Chittenden said.
“Generative AI modeling of single-cell data will eventually afford classification of human disease based on cell differentiation trajectories — how specific cell populations pathologically transform over time to cause disease,” he added.
Such insights into the impact of single cells on disease could shed light on the difference in clinical presentation between a thoracic aortic aneurysm and abdominal aortic aneurysm.
“We have preliminary data indicating abdominal aortic aneurysms associate with additional molecular cell differentiation trajectories [compared to thoracic aortic aneurysms], which are linked to distinct molecular drivers,” Chittenden said. If these findings are supported in humans, drug discovery strategies will include an ability to better understand who is genetically prone to thoracic versus abdominal aortic aneurysms.
The researchers have studied the difference in the two types of aneurysms in mice. “We’re now applying multiple forms of representation learning, such as transfer learning and domain adaptation to see how much of what’s pathologically occurring in mice will inform us on human disease etiology,” Chittenden said.
Moving toward a causal understanding of disease
With appropriate data and analytics, the Genuity Science team’s approach can be applied to almost any phenotype. Using unsupervised machine learning techniques at the single-cell level could lead to more discoveries in the underlying causes of disease, including multiple sclerosis, Parkinson’s disease and non-alcoholic steatohepatitis (NASH).
“It is a very exciting time to be in this space right now — especially at the single cell level,” Chittenden said. A wave of therapeutic breakthroughs could follow from this approach, Chittenden predicted. “It is going to do nothing short of completely transforming our current understanding of human biology and allow us to move collectively forward,” he said.
In particular, such an approach could help scientists move away from understanding not just factors that are correlated with disease, but that cause disease, Chittenden stressed. Although this concept sounds basic, much of the historical focus of genomic research has been on observational data. The interrelated nature of many systems within the body also can make causal understanding of disease elusive.
Another central challenge is scientists’ incomplete understanding of the human genome. “We don’t know what the majority of the genome does yet,” Chittenden said. “Thus limiting ourselves to applications of current knowledge-based AI approaches will not afford true biomedical discoveries.”